Content developed by Kristine Lacek
BLAST (Basic Local Alignment Search Tool)
- Finds regions of similarity between sequences
- Compares a query sequence to a database
- Identifies closest matches
- Uses local alignment
- Finds the best matching regions, not full-length alignment
- Widely used in bioinformatics
- Gene identification
- Species confirmation
- Contamination checks
Different BLAST Programs
blastn- Nucleotide vs nucleotide database
blastp- Protein vs protein database
blastx- Translated nucleotide vs protein database
tblastn- Protein vs translated nucleotide database
megablast- Optimized for highly similar sequences
Databases
- NCBI BLAST
- GISAID BLAST
- Custom Blast Database (CLI)

BLAST Databases Search Considerations
- Databases: NCBI BLAST, GISAID BLAST, Custom Blast Database (CLI)
- NCBI GenBank: open source, INSDC database
- GISAID: access control, but ingests INSDC samples
- For a more complete set of flu samples, which BLAST is better?
- When might an open-source BLAST search be better?

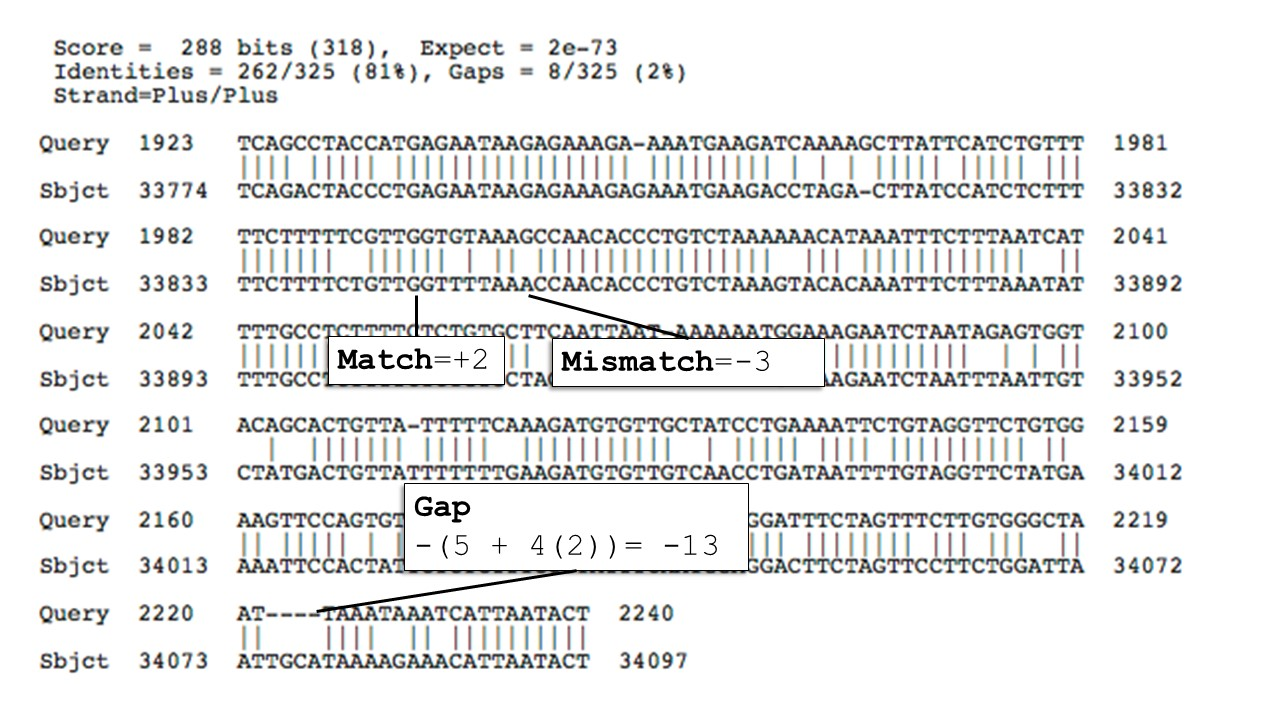
Interpreting BLAST output
- Percent Identity → How similar the aligned region is
- E-value → Probability the match occurred by chance
- Lower = more significant
- Bit Score → Alignment strength (higher = better)
- Alignment Length → How much of the query matched
Running BLAST from the CLI
- BLAST+ is the command-line version
- Installed locally for large datasets or automation
- More reproducible than web BLAST
- Exact parameters and databases can be recorded
- Common workflow
- Prepare database → run BLAST → interpret output
Example command:
blastn -query query.fasta \
-db nt \
-out results.txt
Creating and Using Local Databases
- Download or prepare reference FASTA
- Example: novel influenza genomes for diagnostics efficacy
- Create a BLAST database
makeblastdb -in reference.fasta \ -dbtype nucl - Run BLAST against local database
blastn -query sample.fasta \ -db reference.fasta \ -out results.txt - Faster and ideal for targeted analysis
- Especially useful in viral genomics
Useful CLI Options
- Control output format
-outfmt 6→ tabular format (easy to parse)
- Limit results
-max_target_seqs 5
- Adjust sensitivity
-evalue 1e-5
- Set number of threads
-num_threads 8