Introduction to Bioinformatics Programming
Content developed by Kristine Lacek
Key Points
Bioinformatics programming basics, scripting, and functionality
Presenter Name

Disclaimer
Module Objectives
Key Points
- Trainees will learn scripting basics applicable to any programming language or infrastructure: variables, loops, logic gates, functions, and shebang lines
- Trainees will be able to write simple Bash scripts and customize their shell environment by setting variables, using conditional statements (if/fi), and leveraging loops and aliases for automation and efficiency
- Trainees will understand parallel processing and its utility in optimizing runtime
- Trainees will learn pipe, pipelining, and how it applies to productionalized bioinformatics and ad hoc analyses
Bash Scripting
Key Points
Commands can be run interactively
Enter commands one at a time directly in the terminal
- Commands can be saved in a script file
- A script is a text file (e.g., .sh) containing multiple commands
- Scripts can be executed as a program
- Make executable with
chmod +xscript.sh (permissions!) and run with./script.sh Echo: print text to screen- Shebang line: tells computer how to interpret script
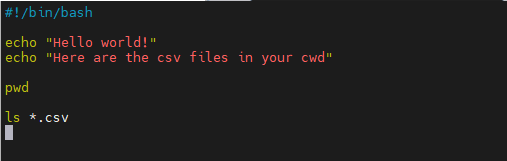
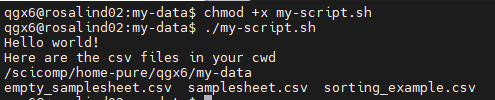
vim
Key Points
Vim is a terminal-based text editor
- Vim is a terminal-based text editor
- Common on servers, HPC systems, and remote machines
- Designed for speed and efficiency
- Keyboard-driven (mouse won’t work!)
- Uses different modes
- Commands behave differently depending on the mode
- Highlights syntax usefully
- Vim demo
- Start and exit
vim file.txt— open a file:wsave:qquit:wqsave & quitq!quit without saving
- Modes
i— insert (edit text)Esc— return to normal mode
- Navigation
- Arrow keys or
h j k l ggtop of fileGbottom of file:<number>go to line number
- Arrow keys or
- Editing
dddelete lineyycopy lineppasteuundo
- Search
/textsearch forwardnnext match

Syntax
Key Points
- Syntax is the set of rules for writing code
- Like grammar in a spoken language
- Strong coffee is good
- El café fuerte es bueno.
- Different programming languages have different syntax
- The same task looks different in Bash, Python, R, etc.
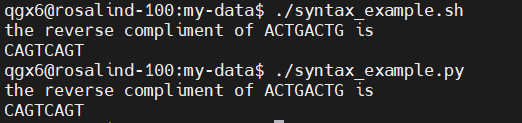
- These two scripts do the same exact task
- Small differences matter
- Symbols, spacing, and keywords must be used correctly
- Using a text editor or Integrative Development Environment (IDE) can help detect errors
- Errors often come from syntax issues
- Missing characters or incorrect formatting can prevent code from running
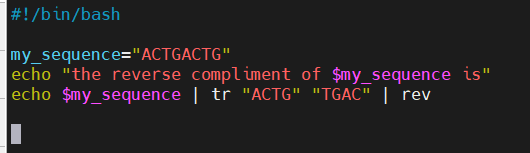
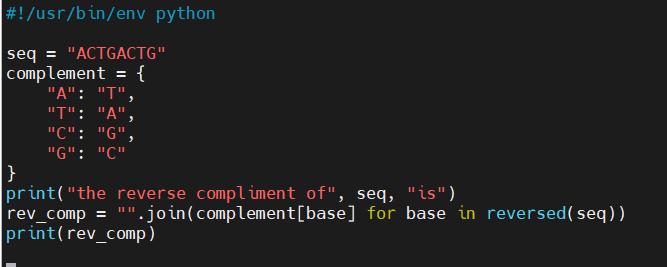
Pseudo Code
Key Points
- Focuses on logic, not syntax
- Describe what the program does without worrying about language rules
- One pseudocode outline can be implemented in Bash, Python, R, etc.
- Helps plan before coding
- Breaks complex problems into clear, manageable steps
- Improves communication
- Easy to read and discuss with collaborators, even non-programmers
Pseudocode for reverse compliment:
Input sequence = “my sequence”
A change to T
T change to A
G change to C
C change to G
Translate nucleotides in “my sequence”
Reverse the translated “my sequence”
Output “my sequence”

Variables
Key Points
- Variables store values like text or numbers
- Commonly used for file names, paths, or sequences
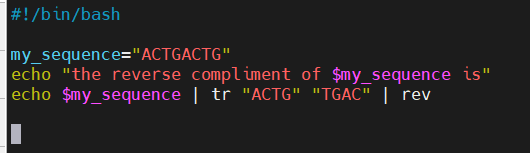
- In syntax example, I set the variable my_sequence to hold “ACTGACTG”
- Assigned without spaces
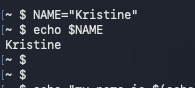
- Accessed with a $
- Good habit to use “$name” to reference
- Useful for making scripts reusable
- Change a variable once instead of editing multiple commands
- Variables can hold many kinds of things:
- Bash variables are untyped by default: everything is treated like a string (not true for every coding language)
- Numbers are handled through context
- Arrays hold multiple values (also called lists in other languages)
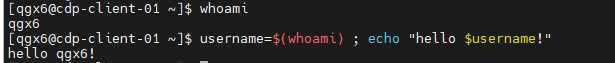
- Variables can even hold the output of another command!
- Subshell
- Current_time as a variable let me run the same code with different results: time changed!
- Choose unique variable names: if you reuse a common variable name, you’ll overwrite it
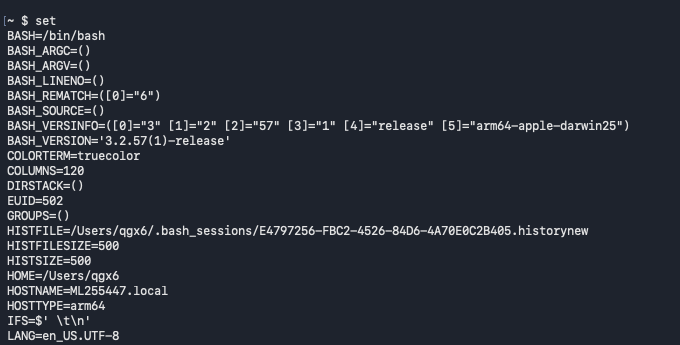
- Certain variables in certain programming languages are already assigned (file, date, etc)
name=value (spaces will cause errors)
Use $name to reference the stored value
VAR=value : Assign a variable
$VAR : Use a variable
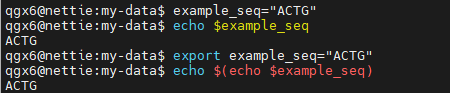
export : Make variable available to subshells
read : Read input from user
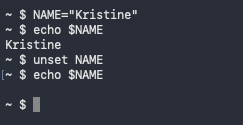
set : Set or unset shell options
unset : Remove variable or function
#!/bin/bash
set –euo pipefail
-e exit on error
-u error on undefined variable
-o pipefail : fail if any command in pipeline fails
Logic: and, or
Key Points
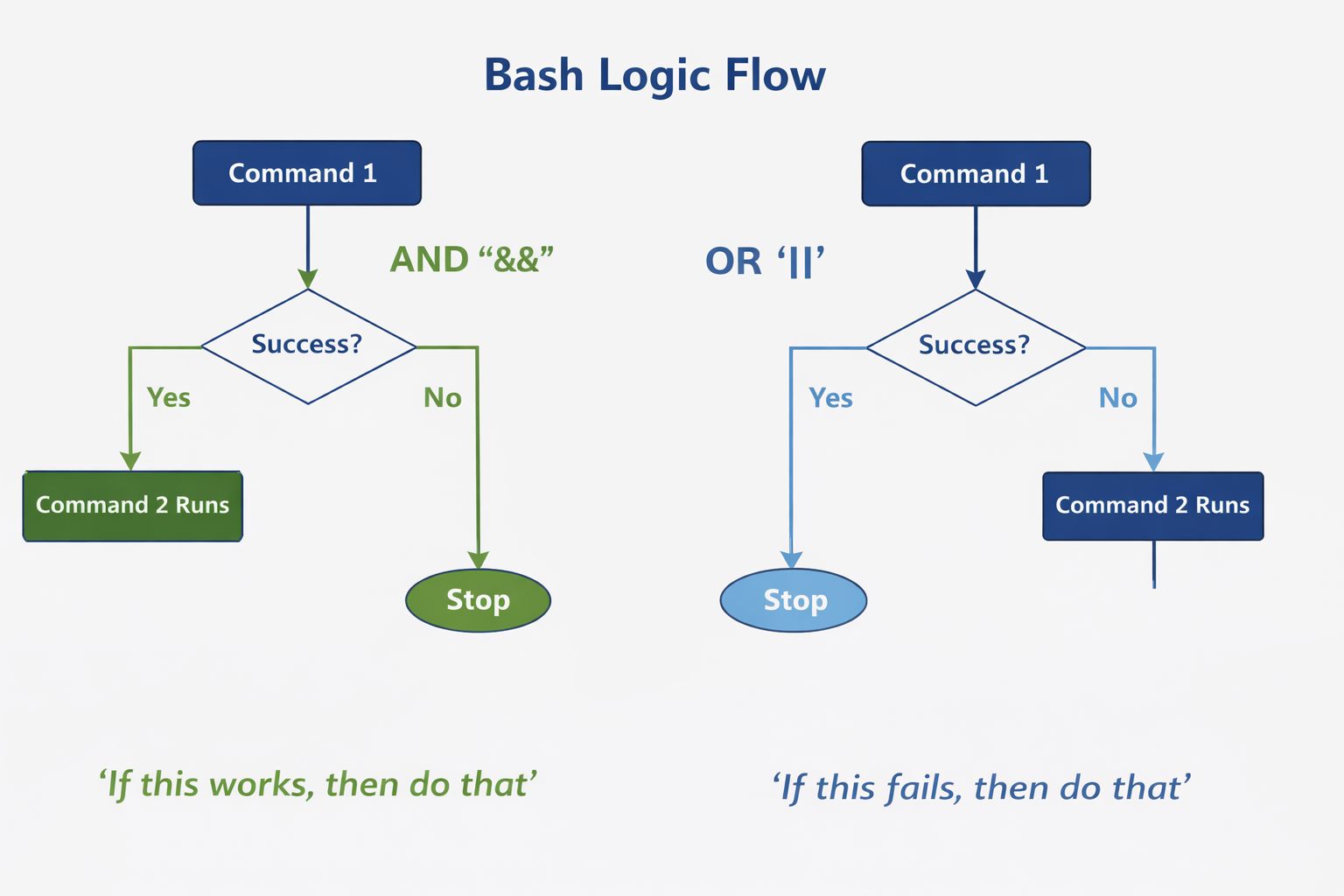
- You can string together multiple commands using and/or logic
Command1 && Command2
Command1 || Command2
- Execute command2 if command1 succeeds
- Execute command 2 if command1 fails, otherwise, do not execute command 2
cd data && echo "Entered data directory" || echo "Could not enter data"


Logic: if, then
Key Points
- Bash if / then Statements
- Used to make decisions in a script
- Run commands only when a condition is true
- Based on command success or a test condition
- Exit status 0 = true, non-zero = false
- Basic structure
if [condition]; then
commands
fi
- Common if conditional flags
- File tests
-e file — path exists
-f file — regular file exists
-d file — directory exists
-s file — file exists and is not empty
- String tests
-z string — string is empty
-n string — string is not empty
string1 = string2 — strings are equal
string1 != string2 — strings are not equal
- Numeric comparisons
-eq — equal
-ne — not equal
-lt — less than
-le — less than or equal
-gt — greater than
-ge — greater than or equal
Logic: else, case
Key Points
- if / else / fi handles yes-or-no decisions
- Run one set of commands or another based on a condition
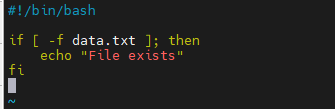
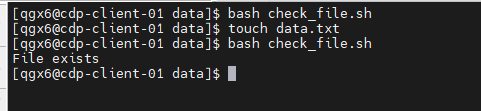
if [ -f data.txt ]; then
echo "File exists"
else
echo "File not found"
fi
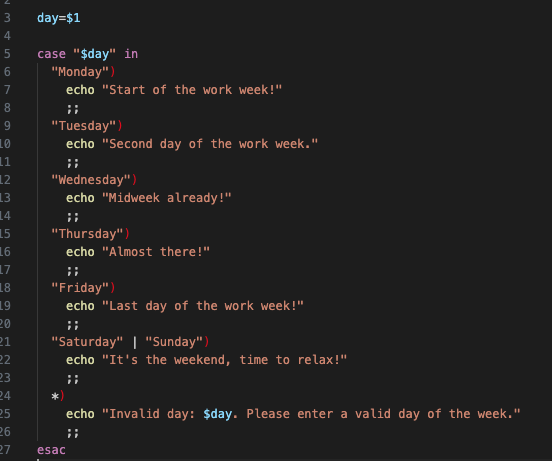
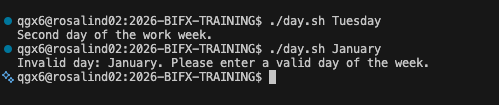
- case / esac handles multiple choices
- Cleaner than many if / else statements
case "$option" in
start) echo "Starting" ;;
stop) echo "Stopping" ;;
*) echo "Unknown option" ;;
esac
Math
Key Points
- Bash does not do math by default
- Arithmetic must be explicitly requested
- Uses integers only by default
- Can use bc –l for floating-point math
- Common operators
+ add
- subtract
* multiply
/ divide
% remainder
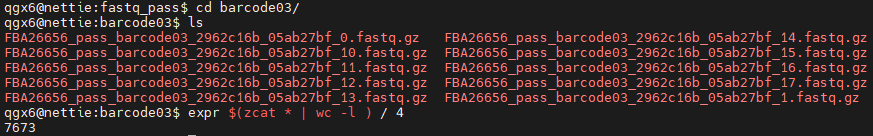
- Math is often used for counters
- Loops, file counts, and simple logic

Loops
Key Points
- Loops repeat commands automatically
- Avoid copying and pasting the same command
- Useful for files, samples, and pipelines
- Common in batch processing and automation
- If you have a nanopore run of 24 samples, you want the same assembly and analysis to happen on each sample
- Run until a condition is met
- Based on lists, counters, or logical tests
- When to use loops
- Process multiple files
- Run the same command on many inputs
- Iterate over values
- Sample IDs, numbers, or directories
- Control workflow logic
- Continue until a condition changes
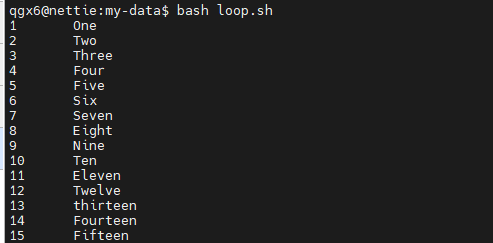
for / do / done
- Loop over a list of values
for file in *.txt; do
echo "$file"
done
while
- Loop while a condition is true
while read line; do
echo "$line"
done < sorting_example.txt
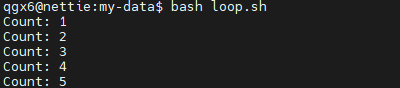
i=1
while (( i <= 5 )); do
echo "Count: $i"
i=$((i + 1))
done
- Loops are powerful but can cause problems
- Infinite loops occur when conditions never change
- Example: while true; do … done
- Forgetting to update loop variables
- Counters or conditions must change inside the loop
- Easy ways to stop a runaway loop
- Ctrl + C to interrupt
- Test with echo before running real commands


Nested Loops
Key Points
- A nested loop is a loop inside another loop
- The inner loop runs completely for each iteration of the outer loop
for i in 1 2 3
do
for j in A B
do
echo “$i $j”
done
done
OUTPUT:
1 A
1 B
2 A
2 B
3 A
3 B
Outer loop runs 3 times
Inner loop runs 2 time for each outer iteration
Total executions: 3x2=6
Functions
Key Points
Functions group related commands
- Package repeated logic into a single unit
- Improve script organization and make long scripts easier to read and maintain
- Defined once, reused many times
- Should have meaningful names to describe what the function does
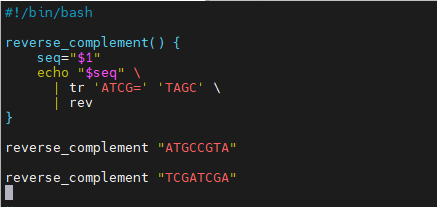
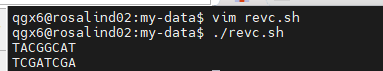
- Function called reverse_complement
$1is the first argument passed to the functiontrperforms base complementation
revreverses the sequence- Function output goes to script output, like a command
- If you notice that you are writing the same code over and over, it can be useful to package that part as a function
- then replace the repeats with said function
- DRY = Don’t Repeat Yourself
- You define the arguments for a function, so it is useful to remember the variables you have already used and make sure not to overwrite them or cause logic errors in the script
- Can define the output of a function as a variable, output to screen, or even a logic evaluation (true/false)

Errors
Key Points
- Common Bash Errors
- Syntax errors
- Missing do, done, then, or fi
- Extra or missing brackets [ ]
- Missing or extra “ “ ( )
- Infinite loops
- Loop condition never changes
- Missing counter update or exit condition
- Permission errors
- Script not executable (chmod +x script.sh)
- No access to files or directories
- Variable mistakes
- Using $var before it is set
- Missing $ when referencing a variable
- A good IDE can help with many of these syntax errors, because it will color code things, make suggestions about what might be common fixes
Debugging Bash scripts
Key Points
- Start with Pseudocode!
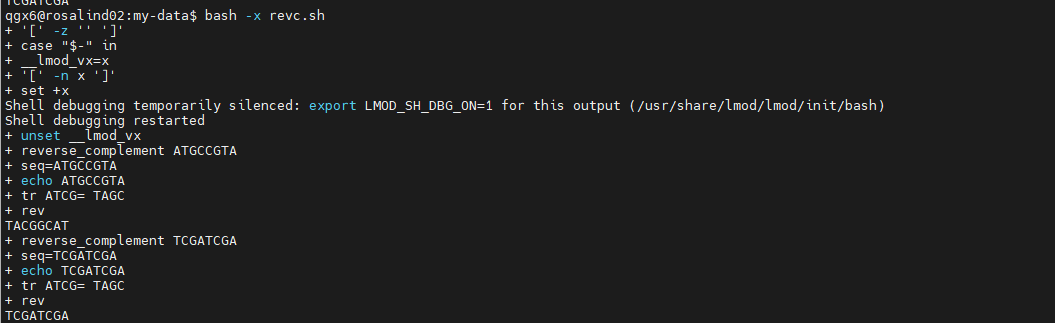
- Run in debug mode
bash -x script.shshows each command as it runs
- Print values to check logic
- Use
echo "$variable"inside loops or conditions - Test commands step by step
- Run lines manually in the terminal
- Start simple and build up
- Confirm each part works before combining commands
Standard Output and Standard Error
Key Points
- Standard Output (stdout)
- Normal program output (results, messages)
- File descriptor 1
- Standard Error (stderr)
- Error, logging, and warning messages
- File descriptor 2
- Allows errors to be handled differently from results
- Useful for scripting and debugging: redirect output and errors independently
- Both go to the terminal by default, but can be redirected separately or together
- Common redirection operators
>redirect stdout2>redirect stderr| tee: writes standard input to standard output
- Useful patterns
- Save results while still seeing errors
- Suppress errors during batch processing
- Example
command > output.txt 2> error.log
Pipelines
Key Points
- Pipelines connect commands together
- Output of one command becomes input to the next
- Use the pipe symbol
| - Pass data through a sequence of tools
- Each tool does one job well
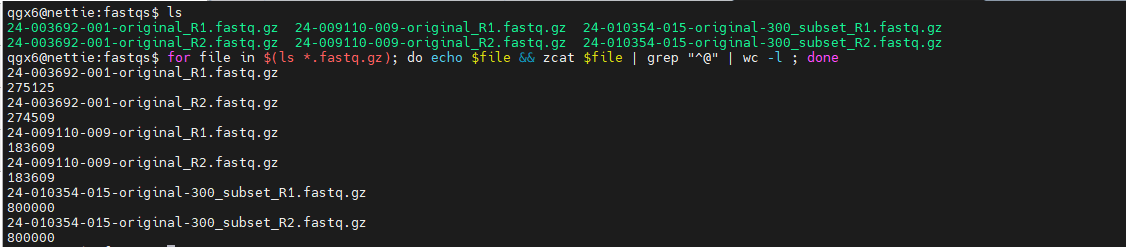
- Common in data processing
- Text files, FASTQ files, and command output
- Bioinformatic pipelines grow from this concept
- Example pseudo code:
#!/bin/bash demultiplex > fastq genome assembly on fastq > fasta update database with genome assembly output curate assembly fastas - Pipelines within pipelines:
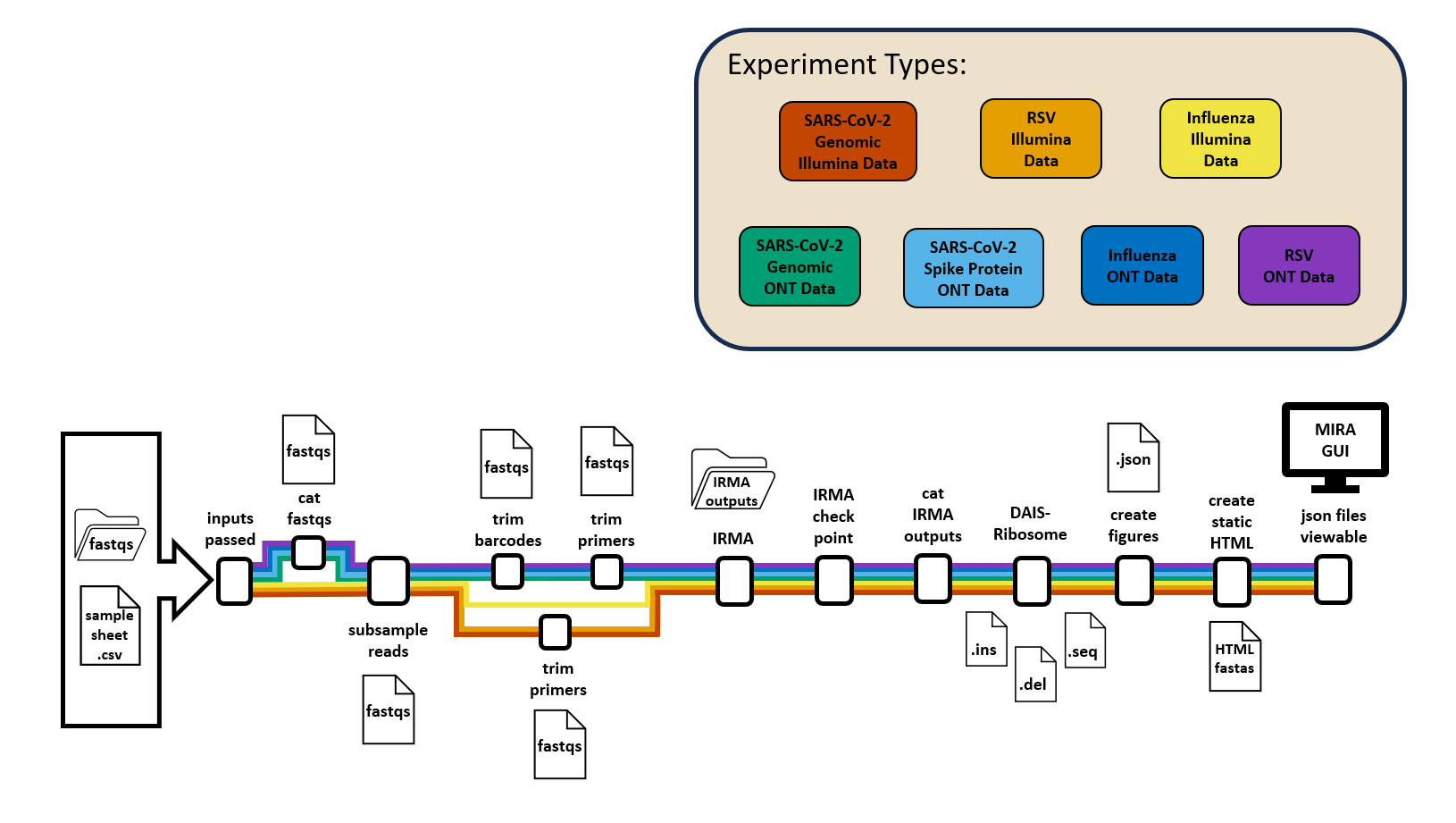
- Genome Assembly: MIRA
check inputs | combine fastqs | subsample reads | trim barcodes | trim primers | run IRMA | check IRMA output | combine output | annotate | create figures
Parallel processing
Key Points
- Two common approaches
- Task parallelism: same command on many inputs
- Data parallelism: split one dataset into parts
- Within tools (multi-threaded software)
- Across tools (running jobs simultaneously)
- Common Bash patterns
- Background jobs (&)
- Job control with wait
- Common bioinformatics tools support multithreading
- Use flags like -t, -p, or –threads
bwa mem -t 8 ref.fa reads.fq > aln.sam
More useful tools and shortcuts
Key Points
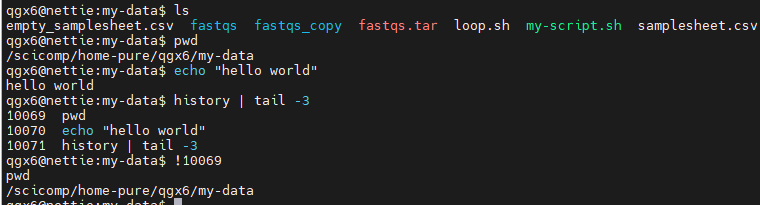
history : Show command history
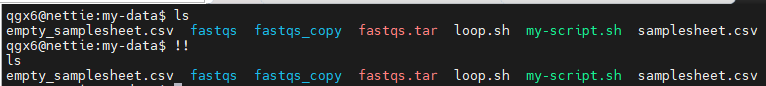
!! : Repeat last command
!n : Run nth command from history
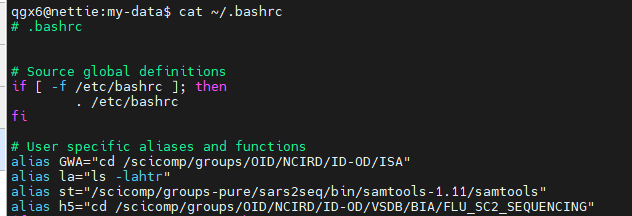
alias : Create command shortcut
unalias : Remove alias
clear : Clear terminal screen
man : Display command manual
help : Show built-in help
date : Show or set system date
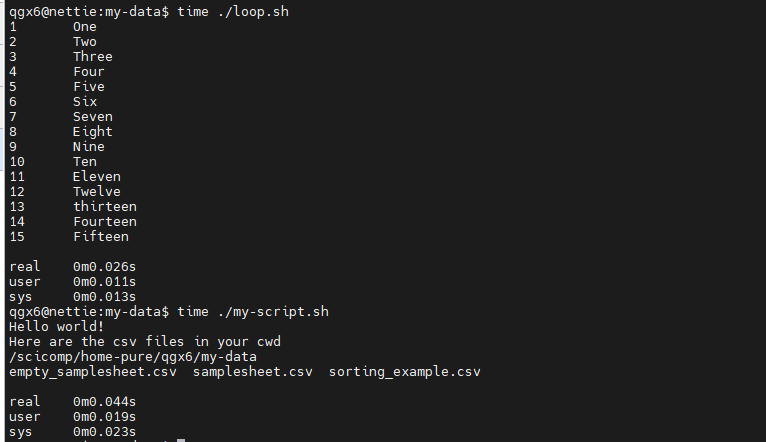
time : Show the runtime of the following command
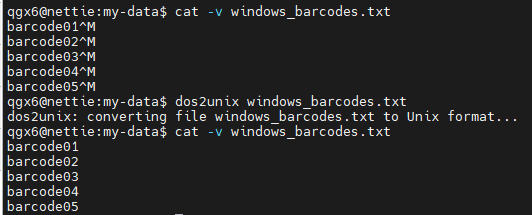
dos2unix : convert Windows file to readable Unix file, remove carriage returns
Bash tip! Ctrl+r to SEARCH your history in command line!
Higher level coding languages
Key Points
- Bash scripting differs from other coding languages
- Abstract away low-level details
- Manage memory, data types, and errors automatically
- Can be compiled into binary
- More expressive and readable
- Fewer lines of code to perform complex tasks
- Rich ecosystems and libraries
- Built-in tools for data analysis, visualization, and networking
Object oriented programming
Key Points
- Group data and actions together
- Similar to treating a file + its operations as one unit
- Classes act like templates
- Define a “sample,” “read set,” or “experiment” once
- Objects represent real things
- Each object holds its own data and methods
- Bash: “Run commands on files”
- Python/R: “Create objects, operate on data”
When to stop using BASH
Key Points
- Logic becomes complex
- Many nested loops, if/else, or long one-liners that are hard to read
- Data structures are needed
- Lists of samples, tables, dictionaries, or metadata don’t fit cleanly in Bash
- Error handling gets messy
- Too many
&&,||, and manual checks for failures - Scripts grow large
- Files longer than ~100–200 lines become difficult to maintain
- You need reproducibility and testing
- Unit tests, versioned packages, and structured logging are easier in Python/R
- Parallelization becomes necessary
- Wrap the bash in a workflow manager like snakemake, wdl, or nextflow
- Visualization is needed
- Better packages and libraries in Python and R
- Other coding languages or packages exist to better handle specific use-cases (biopython, etc)
Packages and tools!
Key Points
- As workflows grow, reuse becomes important
- Copy-pasting scripts leads to errors and drift
- Packages bundle code and functionality
- Install once, use everywhere
- Tools provide standard, tested solutions
- Avoid rewriting common tasks
- Utilize code written by others that is optimized for a certain task
- Easier collaboration and reproducibility
- Others can install the same package and get the same results
- Use Bash to orchestrate tools (in a pipeline)