Quick Example
Here’s a complete working example using test mode (fast configuration for demonstration):
library(nowcastNHSN)
library(dplyr)
# Create test configuration (single state, few draws for speed)
config <- nowcast_config_test(location = "ca", draws = 10)
print(config)
# Fetch data for California
source <- delphi_epidata_source(target = "covid", geo_types = "state")
reporting_data <- fetch_reporting_data(
source = source,
reference_dates = epidatr::epirange(202450, 202510),
report_dates = epidatr::epirange(202450, 202512),
locations = "ca"
)
# Run nowcast - conversion to incremental counts happens internally
results <- run_single_nowcast(
data = reporting_data,
location = "ca",
config = config,
nowcast_date = max(reporting_data$reference_date)
)
head(results, 10)
#> pred_count reference_date draw output_type location
#> 1 723 2024-12-14 1 samples ca
#> 13 723 2024-12-14 2 samples ca
#> 25 723 2024-12-14 3 samples ca
#> 37 723 2024-12-14 4 samples ca
#> 49 723 2024-12-14 5 samples ca
#> 61 723 2024-12-14 6 samples ca
#> 73 723 2024-12-14 7 samples ca
#> 85 723 2024-12-14 8 samples ca
#> 97 723 2024-12-14 9 samples ca
#> 109 723 2024-12-14 10 samples ca
# Summarize to forecast hub format
summary <- summarize_nowcast(results)
head(summary)
#> # A tibble: 6 × 5
#> reference_date location output_type output_type_id value
#> <date> <chr> <chr> <chr> <dbl>
#> 1 2024-12-14 ca quantile 0.01 723
#> 2 2024-12-14 ca quantile 0.025 723
#> 3 2024-12-14 ca quantile 0.05 723
#> 4 2024-12-14 ca quantile 0.1 723
#> 5 2024-12-14 ca quantile 0.15 723
#> 6 2024-12-14 ca quantile 0.2 723
library(ggplot2)
# Pivot quantiles to wide format for ribbon plot
quantile_wide <- summary |>
tidyr::pivot_wider(
names_from = output_type_id,
values_from = value,
names_prefix = "q"
)
nowcast_date <- max(reporting_data$reference_date)
# Get counts at nowcast time (incomplete)
at_nowcast_time <- reporting_data |>
filter(location == "ca", report_date <= nowcast_date) |>
group_by(reference_date) |>
filter(report_date == max(report_date)) |>
ungroup() |>
select(reference_date, at_nowcast = count)
# Get final reported counts (from latest available data)
final_reported <- reporting_data |>
filter(location == "ca") |>
group_by(reference_date) |>
filter(report_date == max(report_date)) |>
ungroup() |>
select(reference_date, final = count)
# Join all data
plot_data <- quantile_wide |>
left_join(at_nowcast_time, by = "reference_date") |>
left_join(final_reported, by = "reference_date")
# Create nowcast visualization
ggplot(plot_data, aes(x = reference_date)) +
# Prediction intervals (draw first so behind)
geom_ribbon(aes(ymin = q0.05, ymax = q0.95, fill = "90% Prediction Interval"), alpha = 0.3) +
# At nowcast time
geom_point(aes(y = at_nowcast, color = "At Nowcast Time"), size = 2) +
geom_line(aes(y = at_nowcast, color = "At Nowcast Time"), linewidth = 1) +
# Nowcast median
geom_point(aes(y = q0.5, color = "Nowcast (Median)"), size = 2, shape = 17) +
geom_line(aes(y = q0.5, color = "Nowcast (Median)"), linewidth = 1, linetype = "dashed") +
# Final reported
geom_point(aes(y = final, color = "Final Reported"), size = 2, shape = 15) +
geom_line(aes(y = final, color = "Final Reported"), linewidth = 1, linetype = "dotted") +
labs(
title = "COVID-19 Nowcast for California",
subtitle = "Comparing nowcast predictions with counts at nowcast time and final values",
x = "Reference Date",
y = "Count",
color = "Count Type",
fill = NULL
) +
theme_minimal() +
theme(
axis.text.x = element_text(angle = 45, hjust = 1),
legend.position = "bottom"
) +
scale_color_manual(values = c(
"At Nowcast Time" = "#E41A1C",
"Nowcast (Median)" = "#377EB8",
"Final Reported" = "#4DAF4A"
)) +
scale_fill_manual(values = c("90% Prediction Interval" = "#377EB8"))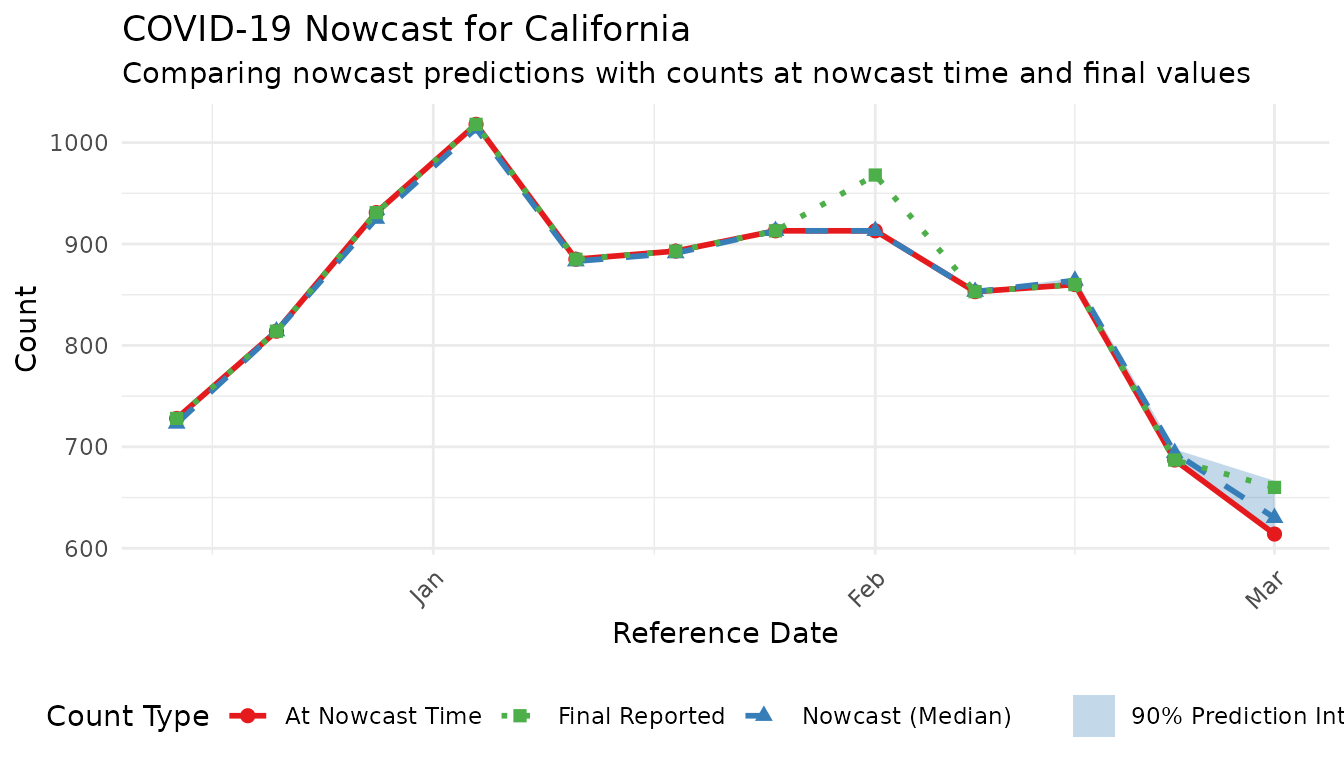
The rest of this vignette provides detailed documentation for production use.
Introduction
This vignette demonstrates how to run NHSN nowcasts for multiple US
states/territories using the nowcastNHSN package. The
workflow supports:
- Batch processing: Run nowcasts for all states in one call
- Single-state execution: Process individual states (useful for Azure task parallelization)
-
Configurable parameters: Customize
max_delay,scale_factor,prop_delay, and uncertainty models -
Partitioned parquet output: Efficient storage and
querying with
arrow
Configuration
The nowcast_config() function creates a validated
configuration object with nowcast parameters:
# Default configuration
config <- nowcast_config()
print(config)
# Custom configuration
config <- nowcast_config(
max_delay = 5,
scale_factor = 3,
prop_delay = 0.5,
uncertainty_model = "skellam", # Options: "negative_binomial", "normal", "skellam"
draws = 500,
output_dir = "output/nowcasts"
)
# Location-specific overrides (optional)
# These override the default config for specific locations
location_configs <- list(
ca = nowcast_config(max_delay = 5, draws = 500),
ny = nowcast_config(max_delay = 7, uncertainty_model = "normal", draws = 500)
)
# Other locations will use the default configConfiguration Parameters
| Parameter | Description | Default |
|---|---|---|
max_delay |
Maximum reporting delay in weeks | 7 |
scale_factor |
Multiplier on max_delay for training volume | 3 |
prop_delay |
Proportion of training data for delay estimation | 0.5 |
uncertainty_model |
Error distribution: "negative_binomial",
"normal", "skellam"
|
"negative_binomial" |
draws |
Number of posterior samples | 1000 |
location |
Single location code (optional) | NULL |
output_dir |
Directory for parquet output | NULL |
Fetching Data
Fetch reporting data for all states in a single API call:
# Create data source
source <- delphi_epidata_source(
target = "covid",
geo_types = "state"
)
# Define date ranges
reference_dates <- epidatr::epirange(202450, 202510)
report_dates <- epidatr::epirange(202450, 202512)
# Fetch all states at once (efficient - single API call)
reporting_data <- fetch_reporting_data(
source = source,
reference_dates = reference_dates,
report_dates = report_dates,
locations = "*" # All locations
)
# Convert cumulative to incremental counts
incremental_data <- cumulative_to_incremental(
reporting_data,
group_cols = c("reference_date", "location", "signal")
)
head(incremental_data)Running Nowcasts
Batch Processing (All States)
# Configure nowcast parameters
config <- nowcast_config(
draws = 100,
output_dir = "output/nowcasts"
)
# Run nowcasts for all states
# This filters the pre-fetched data per state, runs baselinenowcast, and combines results
results <- run_state_nowcasts(
data = incremental_data,
config = config,
locations = "all"
)
# Check for any failures
failed_locations <- attr(results, "failed_locations")
if (length(failed_locations) > 0) {
message("Failed locations: ", paste(failed_locations, collapse = ", "))
}
# View results structure
head(results)Single-State Execution
For Azure task parallelization, you can run individual states:
config <- nowcast_config(
uncertainty_model = "normal",
draws = 100
)
# Run for a single state
ca_results <- run_single_nowcast(
data = incremental_data,
location = "ca",
config = config,
nowcast_date = max(incremental_data$reference_date)
)
head(ca_results)Location-Specific Configurations
Use location_configs to override settings for specific
locations while using a default for the rest:
# Default config for most states
default_config <- nowcast_config(max_delay = 7, draws = 100)
# Overrides for states that need different settings
location_configs <- list(
ca = nowcast_config(max_delay = 5, draws = 100),
ny = nowcast_config(max_delay = 7, uncertainty_model = "normal", draws = 100)
)
results <- run_state_nowcasts(
data = incremental_data,
config = default_config,
location_configs = location_configs,
locations = c("ca", "ny", "tx") # tx uses default_config
)Parallel Processing
Enable parallel processing for faster batch runs:
library(future)
plan(multisession, workers = 4)
config <- nowcast_config(
draws = 100
)
results <- run_state_nowcasts(
data = incremental_data,
config = config,
locations = "all",
parallel = TRUE
)Output
Summarizing Results
Convert raw draws to quantile summaries:
summary <- summarize_nowcast(results)
head(summary)Writing to Parquet
Results are written as partitioned parquet files for efficient querying:
# Write summarized results (default)
write_nowcast_parquet(results, "output/nowcasts")
# Write raw draws instead
write_nowcast_parquet(results, "output/nowcasts_draws", summarize = FALSE)This creates a directory structure like:
output/nowcasts/
├── location=ca/
│ └── part-0.parquet
├── location=ny/
│ └── part-0.parquet
└── location=tx/
└── part-0.parquetReading Parquet Results
library(arrow)
# Read all locations
all_results <- read_nowcast_parquet("output/nowcasts")
# Read specific locations (efficient - only reads needed partitions)
ca_ny <- read_nowcast_parquet("output/nowcasts", locations = c("ca", "ny"))
# Or use arrow directly for more control
ds <- open_dataset("output/nowcasts")
ds |>
filter(location == "ca", reference_date >= "2025-01-01") |>
collect()Command Line Usage
The package includes an entrypoint script for Azure or batch execution:
# Run all states
Rscript inst/scripts/run_nowcast.R --location all --output-dir output/nowcasts
# Run single state (for Azure task)
Rscript inst/scripts/run_nowcast.R --location ca --output-dir output/nowcasts
# Custom parameters
Rscript inst/scripts/run_nowcast.R \
--location ca,ny,tx \
--uncertainty-model skellam \
--max-delay 5 \
--draws 500 \
--output-dir output/nowcasts \
--parallel
# Show help
Rscript inst/scripts/run_nowcast.R --helpVisualizing Results
# Get summary for selected states
summary <- summarize_nowcast(results) |>
filter(location %in% c("ca", "ny", "tx"))
ggplot(summary, aes(x = reference_date)) +
geom_ribbon(aes(ymin = q05, ymax = q95, fill = location), alpha = 0.2) +
geom_ribbon(aes(ymin = q25, ymax = q75, fill = location), alpha = 0.4) +
geom_line(aes(y = median, color = location), linewidth = 1) +
facet_wrap(~location, scales = "free_y", ncol = 1) +
labs(
title = "NHSN COVID-19 Nowcasts by State",
subtitle = "Median with 50% and 90% prediction intervals",
x = "Reference Date",
y = "Predicted Admissions",
color = "State",
fill = "State"
) +
theme_minimal() +
theme(legend.position = "none")Error Handling
The pipeline wraps each state’s nowcast in error handling, allowing the batch to continue even if individual states fail:
results <- run_state_nowcasts(data, config)
# Check which states failed
failed <- attr(results, "failed_locations")
if (length(failed) > 0) {
warning("The following states failed: ", paste(failed, collapse = ", "))
# Results still contain successful states
}Common failure causes: - Insufficient data for a state (not enough reference dates) - Data quality issues (all zeros, excessive missing values) - Numerical issues in uncertainty estimation
Uncertainty Models
The package supports three uncertainty models:
Negative Binomial (Default)
Best for count data with overdispersion. Cannot produce negative predictions.
config <- nowcast_config(uncertainty_model = "negative_binomial")Normal (Rounded)
Symmetric uncertainty around point predictions. Rounds to integers. Can produce negative values for small counts.
config <- nowcast_config(uncertainty_model = "normal")Skellam
Difference of two Poissons. Naturally handles negative revisions. Good for data with frequent downward corrections.
config <- nowcast_config(uncertainty_model = "skellam")