Overview
GISAID, short for the Global Initiative on Sharing All Influenza Data, is an organization that manages a restricted-access database containing genomic sequence data of select virus, primarily influenza viruses. The database has expanded to include the coronavirus responsible for the COVID-19 pandemic as well as other pathogens.
Prerequisites
For all GISAID submissions, seqsender makes use of
GISAID’s Command Line Interface Tools (CLIs) to batch uploading meta-
and sequence-data to their databases. Prior to perform a batch upload to
EpiFlu database, submitters must
- Download the EpiFlu CLI package from the GISAID Platform that is compatible with their machine (e.g., Linux, macOS, or Windows).
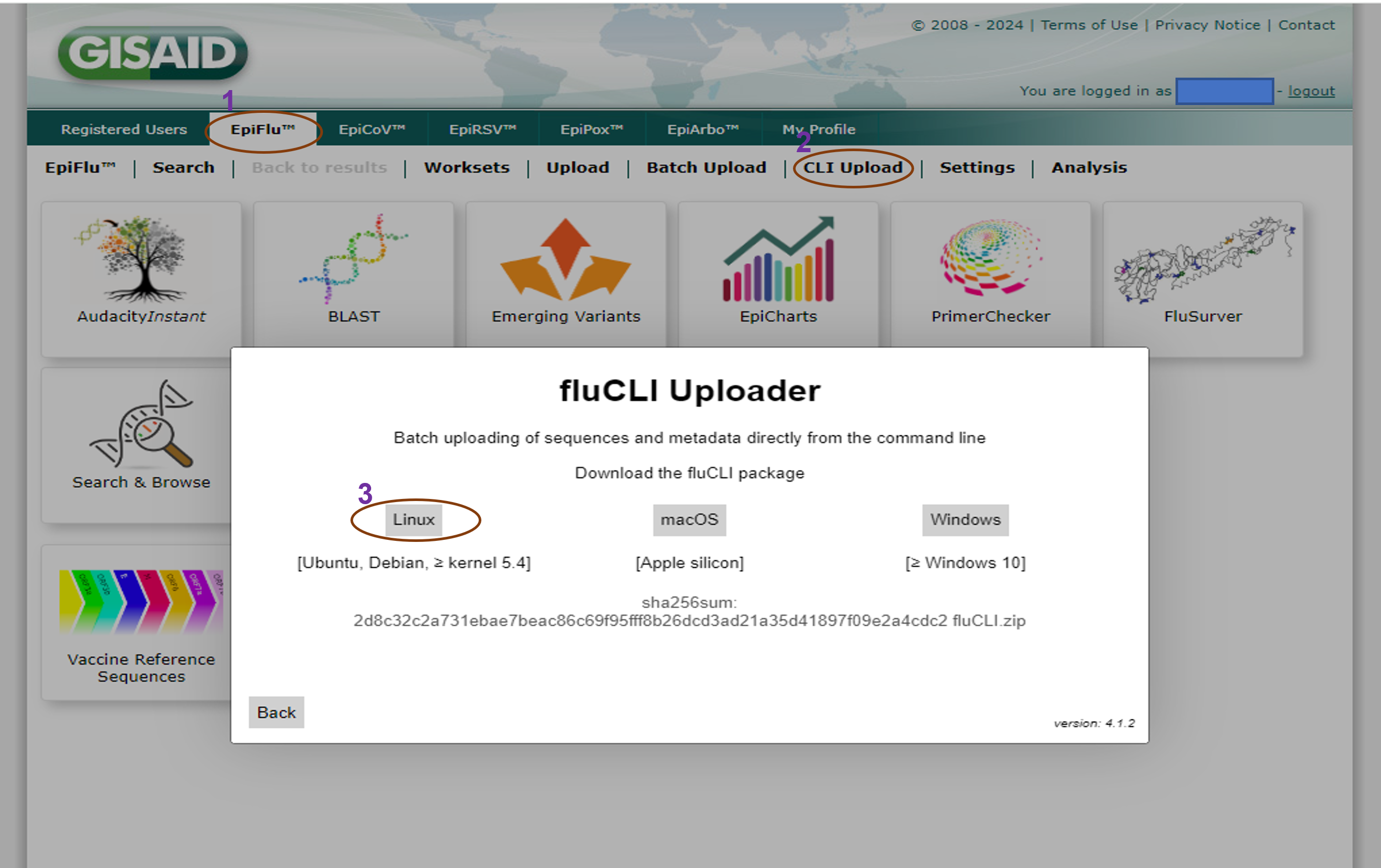
- Unzip the downloaded package and store it in a subfolder called
gisaid_cliwithin a submission directory of choice (e.g.,submission_dir).

Requirement files
After submitters had obtained the GISAID CLI for
EpiFlu, they must also prepare the requirement files
(such as config.yaml, metadata.csv,
sequence.fasta, raw reads, etc.) and store
them in a submission foler of choice (e.g.,
submission_name) within a parent submission directory
(e.g., submission_dir). That way seqsender
will be able to scoop up the necessary files in that folder, generate
submission files, and then batch uploading them to the submitting
database of choices.
Here is a list of the requirement files and where to store them:
-
Config file in a
yamlformat -
Fasta file in a
fastaformat -
Metadata file in a
csvformat

Config file
Config file is a yaml file that provides a brief description about
the submission and contains user credentials that allow
seqsender to authenticate the database prior to upload a
submission.

NOTE:
- To submit to NCBI only, one can remove the
GISAID Submission (b) section from the config file.
Vice versa, to submit to GISAID only, just remove the NCBI
Submission (a) section.
-
Submission_Position determines the order of databases
in which we will submit to first. For instance, if GISAID is set as
1, seqsender will submit to
GISAID first, then after all samples are assigned with a GISAID
accession number, seqsender will proceed to
submit to NCBI. This order of submission ensures samples are linked
correctly between the two databases.
- Username
and Password under the NCBI Submission
(b) section are the credentials used to authenticate the
NCBI FTP Server (not to mistake with individual NCBI
account). See PRE-REQUISITES
for more details.
Fasta file
Fasta file contains nucleotide sequences for all samples. See Genbank Fasta Format for more details.

Metadata file
The metadata worksheet is a comma-delimited (csv) file that contains required attributes that are useful for the rapid analysis and trace back of Influenza A Virus cases.
Here is a short description about the fields in the metadata worksheet.
| Column_name | Description |
|---|---|
| sequence_name | Sequence identifier used in fasta file. This is used to create the fasta file for Genbank or GISAID. |
| organism | The most descriptive organism name for the samples. If relevant, you can search the organism name in the NCBI Taxonomy database. For FLU, organism must be “Influenza A Virus”. For COV, organism must be “Severe acute respiratory syndrome coronavirus 2”. |
| collection_date |
The date on which the sample was collected; must be in the ISO format:
YYYY-MM-DD. For example: 2020-03-25 |
| authors | Citing authors. List of Last, First Middle, suffix separated by a semicolon “;” E.g.: “Baker, Howard Henry, Jr.; Powell, Earl Alexander, III.;” |
| gs-seq_id | Identification to be used for the sequence in the FASTA. |
| gs-Isolate_Name | E.g. “A/Brisbane/1444A/2010” |
| gs-segment | Segment name for GISAID. Options are: “HA”, “HE”, “MP”, “NA”, “NP”, “NS”, “P3”, “PA”, “PB1”, “PB2” |
| gs-Subtype | E.g. “H5N1” |
| gs-Location | E.g., “United Kingdom”, “Japan”, “China”, “United States”, etc. |
| gs-Host | Host or source name., E.g. “human”, “avian”, “chicken”, “Anas Acuta”, “environment”, etc. |
| gs-Collection_Month | For incomplete collection dates, use this field instead of “Collection_Date”. Month of year: “1” = Jan, “2” = Feb, so forth, “12” = Dec |
| gs-Collection_Year | For incomplete collection dates, use this field instead of “Collection_Date”. Four digit year as string: e.g. “2023” |
| gs-Originating_Lab_Id | The numeric ID of the sample”s originating laboratory, e.g. “2698” |
NOTE: The prefix of “gs-” is used to identity attributes for GISAID submissions.
Optional Attributes
To include additional attributes to EpiFlu
submissions, just append gs- in front of the desired
attributes. Here is a list of optional attributes:
| Column_name | Description |
|---|---|
| Lineage | “pdm09”, “Victoria”, “Yamagata” |
| Passage_History | E.g. ‘Original’, ‘E2’,‘MX/C1’, … |
| province | Province of the sample location. |
| sub_province | Sub-Province of the submission location |
| Location_Additional_info | e.g. the city name |
| Host_Additional_info | Any other information about the host or source. |
| Submitting_Sample_Id | Internal ID given to the sample by the submitting lab |
| Originating_Sample_Id | Internal ID given to the sample by the originating lab. |
| Antigen_Character | e.g. ’A/Brisbane/10/2007 |
| Adamantanes_Resistance_geno | “Unknown”, “Resistant”, “Sensitive”, “Inconclusive” |
| Oseltamivir_Resistance_geno | “Unknown”, “Resistant”, “Sensitive”, “Inconclusive” |
| Zanamivir_Resistance_geno | “Unknown”, “Resistant”, “Sensitive”, “Inconclusive” |
| Peramivir_Resistance_geno | “Unknown”, “Resistant”, “Sensitive”, “Inconclusive” |
| Other_Resistance_geno | “Unknown”, “Resistant”, “Sensitive”, “Inconclusive” |
| Adamantanes_Resistance_pheno | “Unknown”, “Resistant”, “Sensitive”, “Inconclusive” |
| Oseltamivir_Resistance_pheno | “Unknown”, “Resistant”, “Sensitive”, “Inconclusive” |
| Zanamivir_Resistance_pheno | “Unknown”, “Resistant”, “Sensitive”, “Inconclusive” |
| Peramivir_Resistance_pheno | “Unknown”, “Resistant”, “Sensitive”, “Inconclusive” |
| Other_Resistance_pheno | “Unknown”, “Resistant”, “Sensitive”, “Inconclusive” |
| Host_Age | Age as string, e.g. “15” |
| Host_Age_Unit | Unit for the age: “Y” for years, “M” for months, “D” for days. |
| Host_Gender | Alternatives are ‘M’ or ‘F’ |
| Health_Status |
For human hosts: Deceased, Recovered, In-patient, Out-patient, Long-term
resident. For animal hosts: Healthy, Sick, Dead. |
| Note | A place for any extra notes about the sample. |
| PMID |
List of PubMed ID’s for referencing PubMed records related to the
sample. Example 1: 1234567; Example 2: 1234567, 1234568 |
You are now ready to install seqsender and batch upload
your submission
Any questions or issues? Please report them on our Github issue tracker.